MICCAI 2014 CLUST is now closed! In this page you can find a summary of the MICCAI workshop, including the final program and the proceedings.
However, we have an open challenge based on the CLUST 2014 data. Results submitted to this challenge will be included in the leaderboard shown below.
Participants agree that the data provided to them shall be used solely for the purpose of participating in the challenge and shall not be used, disclosed or distributed to any other person under any circumstance.
When publishing the results based on the challenge data, participants shall cite the challenge and the references listed at the bottom of this page.
Participants shall submit results and a description of the method, following the instructions listed in the Submission guidelines. The description will serve the organizers to identify and discriminate between methods, but will only be released as informal submission (see below) after explicit request from the participant. The evaluation of the submitted results will be done by the challenge organizers.
By submitting their results, participants agree that these will be included on the online leaderboard. There are two result categories:
Participants shall notify the challenge organizers about any publication which is (partly) based on CLUST data.
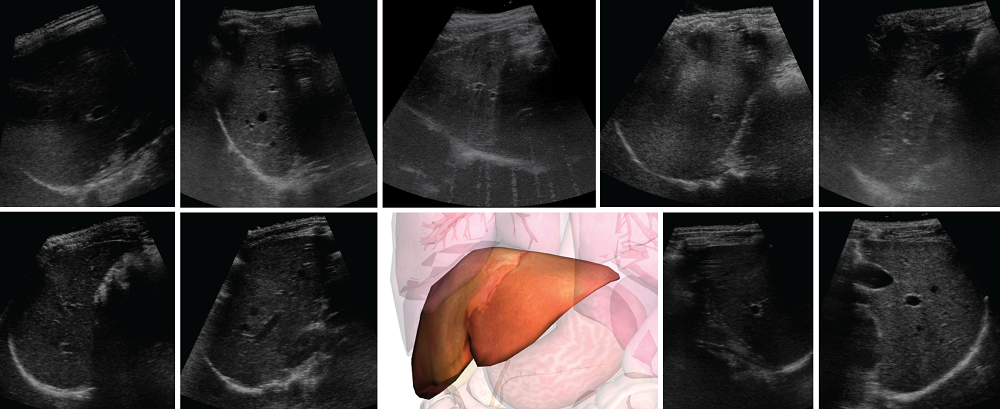
This challenge contains of a total of 54 independent datasets
The data are split into a training (10% of the sequences) and a test set (90%). Annotations are provided for the training set, to allow for some tuning of the tracking algorithm. For the test set, the annotations of the first images are provided. These need to be tracked over time.
The tracking performance of the test set will be evaluated by the organizers and summary results will be provided to the participants and published in Results, as informal or official submission, according to the Submission rules.
For details on the submission formats, check the Submission section in this page.
The training and test data are available here.
A username and password are requested to access the data. Please contact us to receive the login details.
Participants are invited to submit the following files:
It is possible to submit results for all data or for at least one of the following subsets:
Please submit your tracking results in a zip archive to be named as the provided username. Additionally, name the tracking results according to the corresponding sequences and as done for the annotation of the first image. Please submit the tracking results in the same format as provided for the testing sequences of each test set subgroup. In details:
Please include in the archive a text file listing the submitted files and a brief description. Do NOT submit the original images or other tracking results, e.g. deformation fields or tracking results of features which were not provided by the organizers.
The organizers will quantitatively evaluate the submitted tracking results with respect to ground truth annotations. For the evaluation we will consider the following measurements:
We will provide the authors with the outcome of the aforementioned evaluation. After approval of the authors, we will include the quantitative evaluation of the tracking results in Results.
The publication of the results can be of two types:
Participants shall provide a short paper describing their method. Papers should be formatted in Lecture Notes in Computer Science style. The file format for submissions is Adobe Portable Document Format (PDF). Other formats will not be accepted.
Authors of accepted papers shall share their publication, as a reference to the results published in the official Results.
It is possible to upload your submission here.
The results listed on this webpage include all test data (90%) of CLUST14 challenge, while the referenced publications for Sept 2014 state the results for the first test set (70%).
Results of 2D point-landmark tracking. The results are in millimetres and ranked according to increasing mean tracking error.
| Participant | Submission date | |||
|---|---|---|---|---|
| Mean | Standard deviation | 95th percentile | ||
| König L., et al. | 1.51 | 1.88 | 4.06 | Sept 2014 |
| Rothlübbers S., et al. | 1.52 | 1.38 | 4.08 | Sept 2014 |
| Kondo S. | 1.83 | 3.16 | 4.82 | Sept 2014 |
| Benz T., et al. | 1.84 | 2.42 | 5.34 | Sept 2014 |
| Lübke D., and Grozea C. | 1.91 | 2.47 | 5.32 | Sept 2014 |
| Somphone O., et al. | 2.00 | 2.87 | 5.59 | Sept 2014 |
| O'Shea T., et al. | 2.61 | 3.78 | 7.98 | Sept 2014 |
Results of 2D tumor-segmentation tracking. The results are summarized by the range of the mean DICE coefficient, computed for each segmentation and sequence.
| Participant | Submission date | ||
|---|---|---|---|
| Min | Max | ||
| Somphone O., et al. | 76.7 | 92.3 | Sept 2014 |
Results of 3D point-landmark tracking. The results are in millimetres and ranked according to increasing mean tracking error.
| Participant | Submission date | |||
|---|---|---|---|---|
| Mean | Standard deviation | 95th percentile | ||
| Royer L., et al. | 1.62 | 2.19 | 4.81 | Jan 2015 |
| Somphone O., et al. | 2.55 | 2.46 | 7.98 | Sept 2014 |
| Rothlübbers S., et al. | 2.80 | 2.96 | 7.94 | Sept 2014 |
| Lübke D., and Grozea C. | 4.63 | 4.03 | 12.44 | Sept 2014 |
All deadlines are 23:59 Pacific Standard Time.
| Mar 1 | Workshop acceptance notification | |
| Mar 25 | Release of 80% of the data | |
| Mar 30 | Call for papers | |
| Jun 18 | Tracking results submission deadline | |
| Jun 19 | Tracking performance notification | |
| Jun 24 | Paper submission deadline | |
| Jul 5 | Acceptance/rejection notification | |
| Jul 22 | Submission deadline for accepted papers | |
| Jul 22 | Workshop registration due | |
| Aug 1 | Final program | |
| Sep 8 | Release of final 20% of the data | |
| Sep 14 - PM | Workshop date |
Download the detailed schedule here.
| 1.15 - 1.40 pm | Welcome and Introduction C. Tanner, V. De Luca, ETH Zurich, Switzerland |
| 1.40 - 3.00 pm | Oral Session 1 - 2D Tracking Chair: A. Cifor, University of Oxford, UK |
| Liver Feature Motion Estimation in Long High Frame Rate 2D Ultrasound Sequences. T. O'Shea, J. Bamber, E. Harris, ICR, London, UK | |
| Liver Ultrasound Tracking Using Long-term and Short-term Template Matching. S. Kondo, Konica Minolta Inc., Osaka, Japan | |
| Kernel-based Tracking in Ultrasound Sequences of Liver. T. Benz1, M. Kowarschik1,2, N. Navab1, 1 CAMP TU Munich, Germany; 2 Angiography & Interventional X-Ray Systems, Siemens AG, Healthcare Sector, Forchheim, Germany | |
| A Non-Linear Image Registration Scheme for Real-Time Liver Ultrasound Tracking using Normalized Gradient Fields. L. König, T. Kipshagen, J. Rühaak, Fraunhofer MEVIS, Lübeck, Germany | |
| 3.00 - 3.30 pm | Coffee Break |
| 3.30 - 4.30 pm | Oral Session 2 - 2D and 3D Tracking Chair: M. A. Lediju Bell, Johns Hopkins University, Baltimore, USA |
| High Performance Online Motion Tracking in Abdominal Ultrasound Imaging. D. Lübke, C. Grozea*, Fraunhofer MEVIS, Bremen, Germany; * Fraunhofer FOKUS, Berlin, Germany | |
| MICCAI CLUST 2014 - Bayesian Real-Time Liver Feature Ultrasound Tracking. S. Rothlübbers, J. Schwaab*, J. Jenne, M. Günther, Fraunhofer MEVIS, Bremen, Germany; *Mediri GmbH, Heidelberg, Germany | |
| Live Feature Tracking in Ultrasound Liver Sequences with Sparse Demons. O. Somphone, S. Allaire, B. Mory, C. Dufour, Medisys Lab Philips Research, Suresnes, France | |
| 4.30 - 5.00 pm | Overall Results and Conclusions V. De Luca, ETH Zurich, Switzerland |
The workshop was held in the Stata Center 36, in room 428. A map of the MIT campus can be found here.
Further information can be found on the MICCAI Workshop page.
The proceeding of MICCAI CLUST 2014 are available for download here.
A summary of the results is published in Results, see Submission date: Sept 2014. These results are obtained from all test data. In the proceedings, authors reported tracking results only for the initial test data (70% of all data).
Valeria De Luca, ETH Zurich, Switzerland vdeluca@vision.ee.ethz.ch
Amalia Cifor, University of Oxford, UK amalia.cifor@eng.ox.ac.uk
Muyinatu A. Lediju Bell, Johns Hopkins University, USA muyinatu.ledijubell@jhu.edu
Christine Tanner, ETH Zurich, Switzerland tannerch@vision.ee.ethz.ch